|
8/13/2023 0 Comments Designing primers using amplifx
Nucleic Acids Res 29:2850–2859īlack DL (2003) Mechanisms of alternative pre-messenger RNA splicing. Modrek B, Resch A, Grasso C et al (2001) Genome-wide detection of alternative splicing in expressed sequences of human genes. 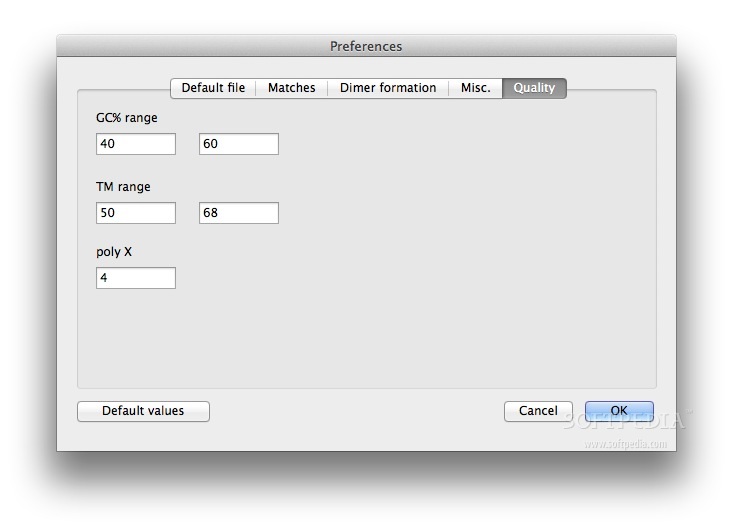
Nilsen TW, Graveley BR (2010) Expansion of the eukaryotic proteome by alternative splicing. Pan Q, Shai O, Lee LJ et al (2008) Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Wang ET, Sandberg R, Luo S et al (2008) Alternative isoform regulation in human tissue transcriptomes. Samuels DC, Han L, Li J et al (2013) Finding the lost treasures in exome sequencing data. Venter JC, Adams MD, Myers EW et al (2001) The sequence of the human genome. Quackenbush J (2001) The power of public access: the human genome project and the scientific process. In this chapter we describe a protocol to efficiently detect, analyze, and quantify alternative splicing patterns of immune mediators such as chemokines, cytokine and their receptors and ligands in cancer by quantitative PCR. In order to understand the role that such splice variants play in cancer, it is vital to be able to accurately quantify their expression levels in different cell types and organs, both in normal conditions and in disease. Therefore, splice variants are critical tools to assess disease progression and clinical prognosis, and hold great promise as potential targets for therapeutic intervention. Moreover, changes in the relative amounts of different splice variants derived from the same pre-mRNA are a hallmark in cancer, and aberrant expression of alternatively spliced mRNAs has been linked to cancer initiation and progression. 
Alternative splicing evolved as a very efficient way to generate proteome diversity and to regulate cell homeostasis from a limited number of genes.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
